Researchers Discover Wide Variation in Virulence of Non-Pathogenic Fungi
By Andy Flick, Evolutionary Studies scientific coordinator
A new study led by research assistant professor David Rinker sheds light on how fungal pathogenicity might evolve. The article, ŌĆ£ was published in the journal Communications Biology in September 2024.
According to Rinker, ŌĆ£different isolates or strains of Aspergillus fumigatus have already been shown to vary widely at the genomic level and some isolates of A. fumigatus are known to have greater pathogenic potentials than others.ŌĆØ The group set out to test if that wide range of virulence could also be found in the closely related non-pathogenic species Aspergillus fischeri.
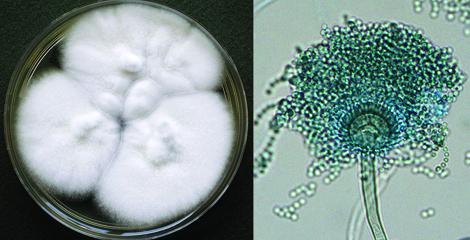
In this study, mice and mouse immune cells were infected with spores from many different strains of Aspergillus fischeri. By measuring mortality – or the ability for the fungus to kill its host or host cells, they were able to assess how each strain affected host survival. While infections by some strains had no impact, others significantly affected mouse survival. Thus, some strains appear to be non-pathogenic while others are highly pathogenic.
The studyŌĆÖs findings underscore the need for a broader perspective on fungal virulence that includes non-pathogenic species, which may harbor hidden pathogenic potential that could emerge under certain environmental conditions or in immunocompromised hosts.
ŌĆ£As far as we know,ŌĆØ Rokas said, ŌĆ£this is the first published study to show this variability in a non-pathogenic species of fungus. When we give research talks on our findings, people get really excited because our results suggest that we need to start thinking of these traits like virulence, not as a single point from a reference strain, but more like a distribution. ThatŌĆÖs the really cool result!ŌĆØ
Rinker noted that, ŌĆ£fungal pathogenicity is not considered obligate, but rather opportunistic. Therefore, the potential for new, opportunistic fungal pathogens to make the ŌĆśleapŌĆÖ into the clinic may be greater than we previously realized.ŌĆØ He continued, ŌĆ£looking for shared mechanisms of virulence in ŌĆśnon-pathogensŌĆÖ may shed light into how pathogens originate or even predict the emergence of novel pathogens.ŌĆØ
Citation: Rinker, D.C., Sauters, T.J.C., Steffen, K. et al. Strain heterogeneity in a non-pathogenic Aspergillus fungus highlights factors associated with virulence. Communication Biology 7, 1082 (2024). https://doi.org/10.1038/s42003-024-06756-8
Funding: Computational infrastructure at ╣·▓·┤½├Į was provided by The Advanced Computing Center for Research and Education (ACCRE). This research was supported by the National Institutes of Health National Institute of Allergy and Infectious Diseases (R01 AI153356). Research in the Rokas Lab is also supported by the National Science Foundation (DEB-2110404) and the Burroughs Wellcome Fund. Research in the Goldman lab is also supported by the Funda├¦├Żo de Amparo ├Ā Pesquisa do Estado de S├Żo Paulo (FAPESP) grant numbers 2021/04977-5 and 2023/00206-0 (ED), the Conselho Nacional de Desenvolvimento Cient├Łfico e Tecnol├│gico (CNPq) grant numbers 301058/2019-9 and 404735/2018-5, and the CNPq, FAPESP and Funda├¦├Żo Coordena├¦├Żo de Aperfei├¦oamento do Pessoal do Ensino Superior (CAPES) grant number 405934/2022-0 (The National Institute of Science and Technology INCT Funvir), all from Brazil. GoldmanŌĆÖs lab was also funded by the Joint Canada-Israel Health Research Program, jointly supported by the Azrieli Foundation, CanadaŌĆÖs International Development Research Center, Canadian Institutes of Health Research, and the Israel Science Foundation.